UK Biobank data is an invaluable resource for genetic research, containing genetic, lifestyle, and health information from half a million individuals. To leverage this wealth of data, DISGENET v25.1 features the integration of a dataset that provides genome-wide associations for Electronic Health Record (EHR)-derived phenotypes from White British participants in the UK Biobank (data available at PheWeb UKB-TOPMed).
However, while integrating these data, a significant challenge emerged: aligning the 1,419 phenotypes encoded using Phecodes with the DISGENET coding system, which relies on Unified Medical Language System (UMLS) Concept Unique Identifiers (CUIs).
What are Phecodes?
When conducting a genome-wide association study (GWAS) using UK Biobank data, one of the key challenges is harmonizing phenotype definitions. Phecodes help address the harmonization of different phenotype definitions by grouping ICD (International Classification of Diseases) codes into clinically meaningful categories, enhancing the statistical power of both GWAS and Phenome-Wide Association Studies (PheWAS).
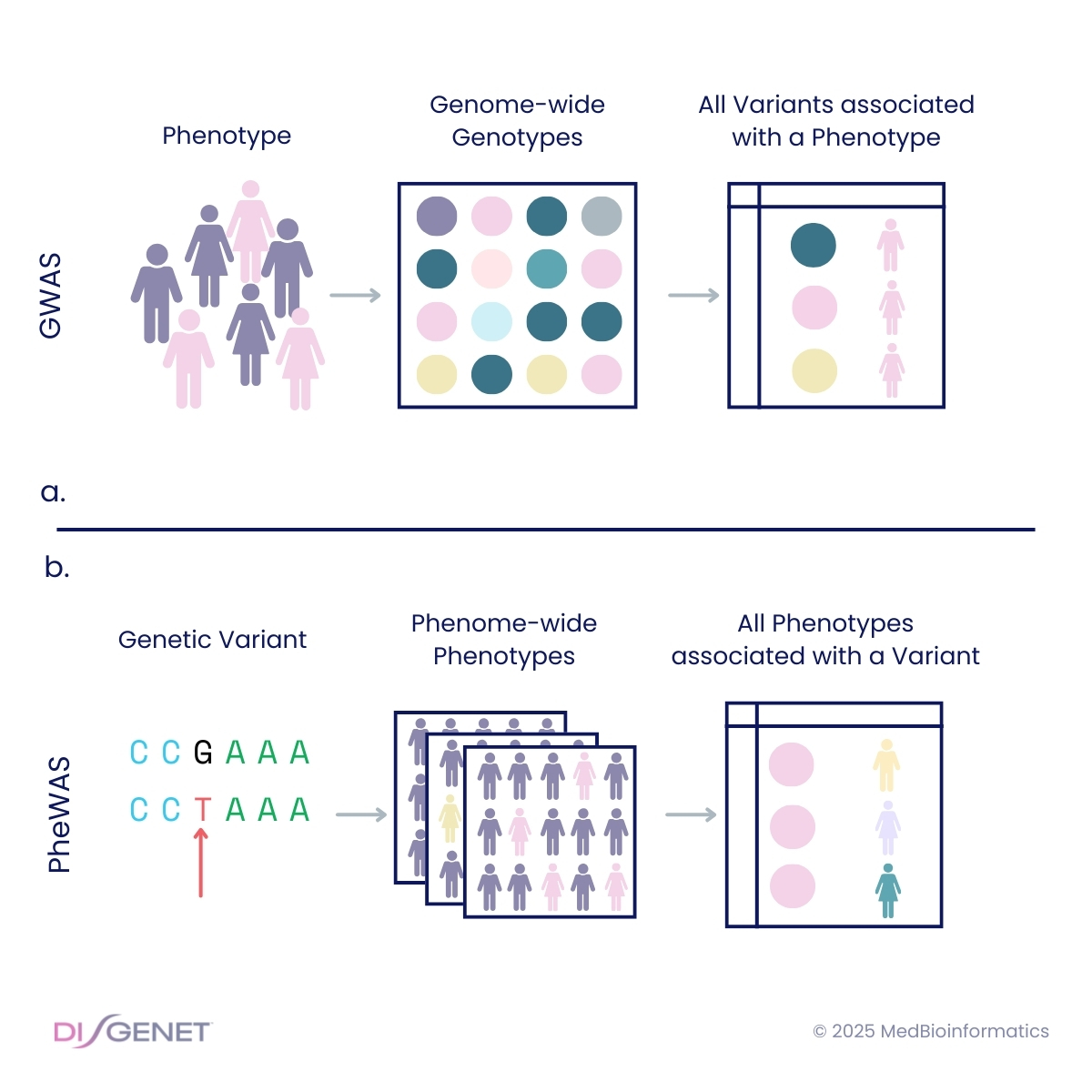
Initially developed for Phenome-Wide Association Studies to facilitate the exploration of genetic associations across multiple phenotypes, Phecodes now play a crucial role in EHR phenotyping. Their ability to aggregate similar conditions into broader disease categories simplifies large-scale genetic analyses and prevents phenotypic data fragmentation.
Phecodes are maintained by the Center for Precision Medicine at Vanderbilt University Medical Center and are available in the PheWAS Catalog. The latest version (1.2) includes 1,867 Phecodes, derived from approximately 15,500 ICD-9-CM codes and 90,000 ICD-10-CM codes.
Challenges in Mapping Phecodes to UMLS CUIs
Phecodes are hierarchical, capturing disease phenotypes at multiple levels:
- Three-digit Phecodes (e.g., 301 – Personality disorders) represent broad phenotypes.
- More specific Phecodes (e.g., 301.1 – Affective personality disorder, 301.12 – Chronic depressive personality disorder) refine the classification.
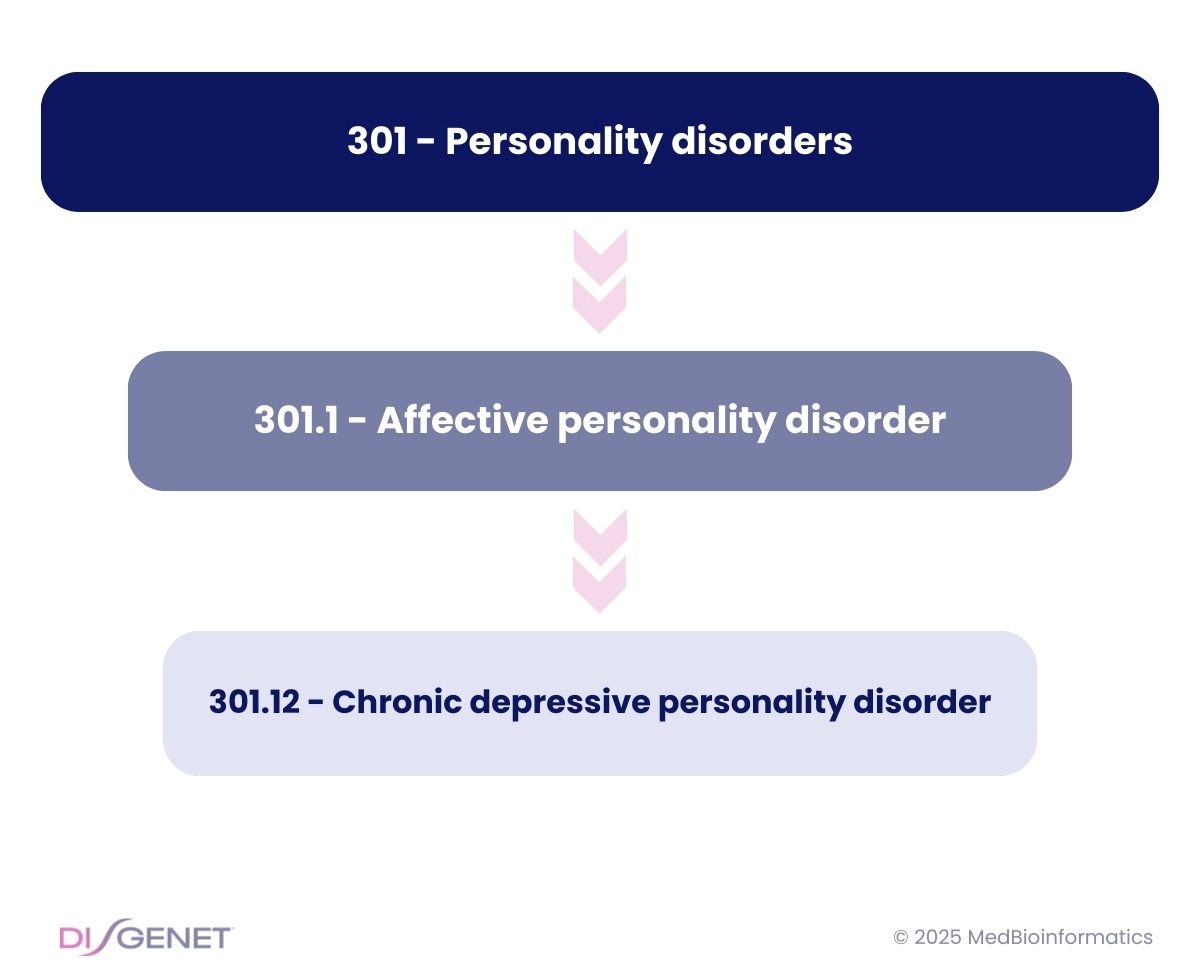

A single Phecode often corresponds to one or several ICD codes. However, in some cases, a single Phecode can represent over 100 different ICD codes. This wide variation presents a significant challenge when combining Phecode-based data with datasets featuring other clinical coding systems or disease ontologies.
Because of this multiplicity, to integrate this dataset with other data in DISGENET, coded using UMLS CUIs, we decided to find a way of mapping the Phecodes to one or a few concepts that best capture each Phecode. The alternative approach of mapping Phecodes to all their corresponding ICD codes and subsequently using UMLS cross-references to retrieve the CUIs would result in redundant representations of genetic associations within DISGENET, potentially inflating the data and introducing biases in downstream analyses.
Our Approach: Mapping Phecodes to CUIs
Rather than converting Phecodes to ICDs first and then mapping to CUIs, we developed a direct mapping strategy to associate each Phecode with one or two CUIs that best capture its meaning. UMLS CUIs serve as standardized references within the UMLS, enabling integration across multiple biomedical datasets and ensuring uniformity in disease representation.
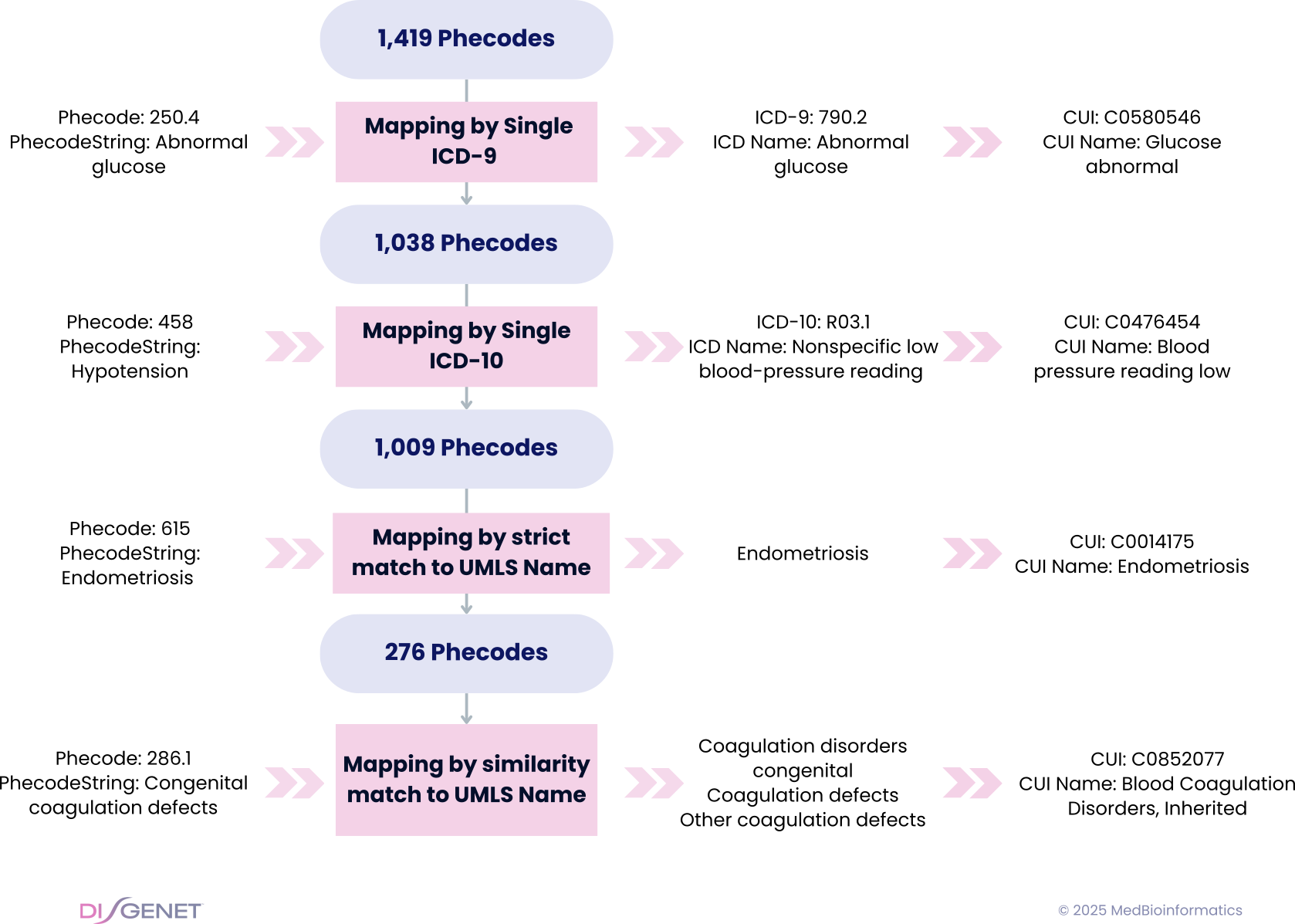
Our 4-step strategy was as follows:
- 1. Mapping by Single ICD-9: If a Phecode maps exclusively to one ICD-9 code, we use it to map the Phecode to a UMLS CUI via UMLS cross-references.
- 2. Mapping by Single ICD-10: If a Phecode maps exclusively to one ICD-10 code, we use it to map the Phecode to a UMLS CUI via UMLS cross-references.
- 3. Strict Matching by Phecode Name: If a Phecode maps to multiple ICD codes, we look for an exact match between the Phecode name (PhecodeString) and a CUI name from the MRCONSO (UMLS Meta-thesaurus Concept Names and Sources) file. If multiple matches exist, we keep the CUI associated with the largest number of genes in DISGENET.
- 4. Similarity-Based Matching by Phecode Name: When multiple ICD codes are linked to a Phecode, and no concept name matches exactly the PhecodeString, we apply a string similarity-based algorithm to find the best match from the UMLS MRCONSO concept names, followed by expert review for lower-confidence cases.
Key Takeaways from Phecode Mapping
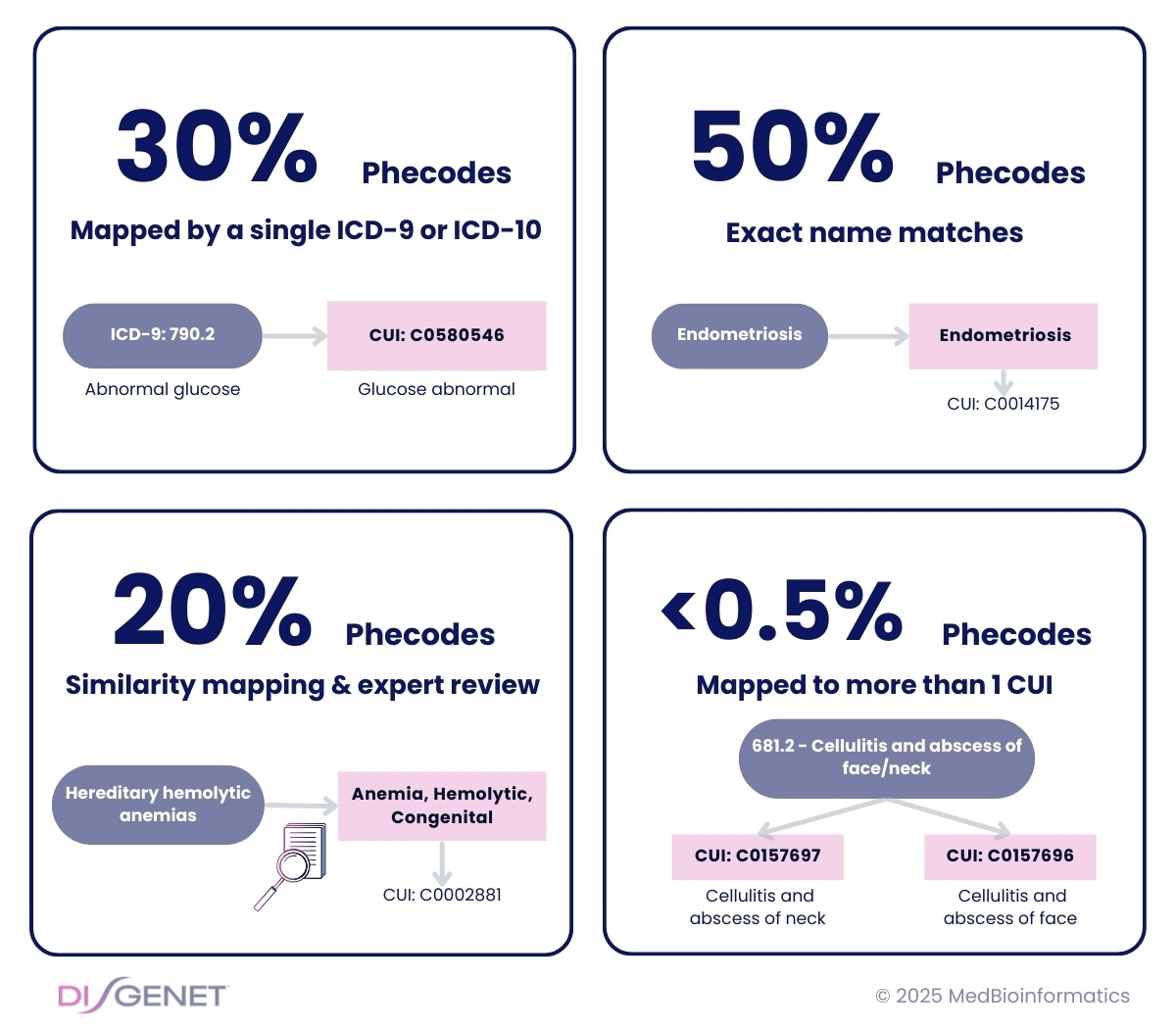
Single ICD Code Mapping
30% of Phecodes had a single ICD-9 or ICD-10 code which we used to map to CUIs.
Exact Name Matches
Over 50% of Phecodes had names that exactly matched one CUI.
Similar Names Matches
Expert curation was necessary to identify the more appropriate CUI for 20% of Phecodes.
Multiple CUIs for Some Phecodes
Some Phecodes mapped to multiple CUIs. In these cases, we keep the one with information in the DISGENET database.
Expert Review Ensured Accuracy
After expert review of the mappings using the similarity-based approach, all Phecodes could be mapped to CUIs.
Mapping to Two CUIs
In the final dataset, only 5 Phecodes required mapping to two CUIs, mainly due to conditions affecting different anatomical sites (e.g., 681.2 – Cellulitis and abscess of face/neck → C0157696; C0157697, Cellulitis and abscess of face; Cellulitis and abscess of neck).
Complex Mapping for One Phecode
Only one phecode (225.1 – Benign neoplasm of brain, cranial nerves, meninges) mapped to three CUIs due to the same reason (C0004992, C0154033, C0496899. Benign neoplasm of cranial nerves; Benign neoplasm of cerebral meninges; Benign neoplasm of brain).
Challenges in Mapping Phecodes for GWAS
Need for Expert Review
Expert review was necessary to ensure accurate and less redundant mapping.
Limitations of Large Language Models (LLMs)
Automating this process with Large Language Models (LLMs) was not feasible, as they perform poorly in entity normalization.
Granularity of UMLS CUIs
UMLS CUIs offer the granularity needed for this task, as they support both specific disease terms and broader hierarchical navigation, aligning well with phecode structures.
Advancing GWAS Integration in DISGENET
This Phecode-to-CUI mapping will enable researchers working with UK Biobank GWAS data to align their findings with other datasets integrated into DISGENET, enhancing the utility and interpretability of genetic discoveries.
By directly linking Phecodes to CUIs, we improve disease concept harmonization, ensuring that GWAS insights can be integrated with broader biomedical knowledge resources.
Notably, the implemented mapping procedure is designed for reusability, enabling the integration of diverse EHR-derived phenotype datasets.
