DISGENET, the largest and most extensive gene-disease association network, has released version 24.3, featuring several new additions.
DISGENET version 24.3 includes:
- The DISGENET AI Assistant (Beta)
- New Clinical Trial Data Source
- Expanded Data Coverage
- Cytoscape App Improvements
- Enhanced REST API Filtering
With these enhancements, DISGENET continues to provide you with reliable sources, accessible data, and actionable tools to support your research all within one platform.
The DISGENET AI Assistant
We have released a beta version of our new tool, the DISGENET AI assistant, to simplify the user’s interaction with the platform. It combines the power of generative AI with the capabilities of the DISGENET REST API. Ask questions in natural language, and the assistant will translate them into structured queries to the database, leveraging all of DISGENET’s features.
In addition, it summarizes the main results of the queries, helping researchers gain a deeper understanding of the data by providing context and explanations.
As a beta version, your feedback is crucial in helping us refine and improve this tool. Please share your thoughts and experiences with us.

The DISGENET AI Assistant is available on Standard & Advanced plans. Learn more about our plans here or try it out by registering for a free 7-day trial.
New Clinical Trial Data Source
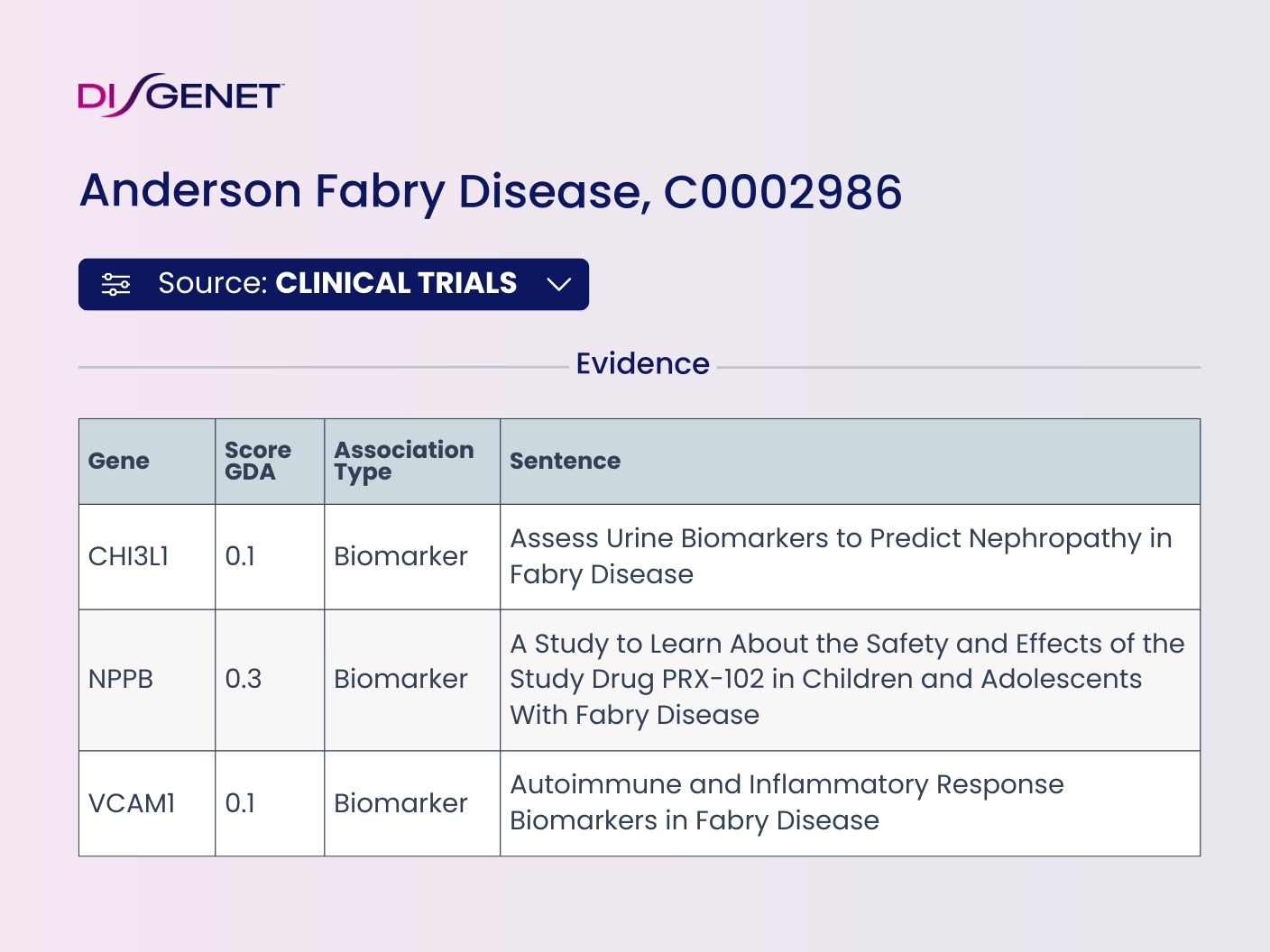
We identify biomarkers of different types in clinical studies from ClinicalTrials.gov using our NLP tools. Clinical trial data, particularly the biomarkers measured within these trials, offers significant value when integrated with other DISGENET data.
By combining clinical trial data with preclinical data from literature and databases, researchers can gain deeper insights into disease mechanisms, identify potential biomarkers, validate existing ones, evaluate drug efficacy, and explore new drug applications.
Data on tables can be filtered by the source Clinical Trials on the web. Use the Source button above the table view to select data specifically from this source. The API calls can also be filtered by this new datasource.
Expanded Data Coverage
DISGENET v24.3 offers a significant expansion of its data coverage, providing researchers with a wealth of information.
Here’s a breakdown of the key metrics:
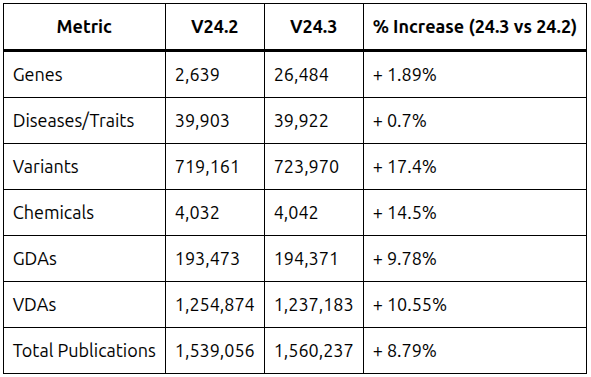
DISGENET’s expanded data coverage comes from trusted sources, including:
- Curated Databases: CLINGEN, ClinVar, MGD (human), Orphanet, PsyGeNET, RGD (human), UniProt
- Clinical Trial Data (new in v24.3)
- Inferred Data: HPO, GWAS Catalog, PheWAS Catalog
- Model Organisms: MGD (Mouse), RGD (Rat), text mining (several models)
- Text Mining: from our high-performance NLP tool
DISGENET Cytoscape App Improvements
The DISGENET Cytoscape App (v 8.0.3) has been significantly upgraded, providing you with enhanced capabilities for network analysis.
The key Cytoscape app improvements are:
Search for up to 100 entities at once
Conduct more comprehensive searches with up to 100 comma-separated entities (genes, variants, and diseases) in a single query. This enables you to efficiently investigate more complex networks without the need for multiple, time-consuming queries.
Use NCBI gene identifiers
NCBI gene identifiers are widely used and now you can query the DISGENET database directly using them on the Cytoscape App.
Improved search box suggestions
The app now provides more relevant suggestions as you type in the entity search boxes, helping you to find the right genes, variants, and diseases more quickly and easily.
Enhanced REST API Filtering
DISGENET v24.3 introduces new filtering and sorting options for the GDAs and VDAs endpoints, providing you with greater flexibility and control over your data retrieval.
New options include:
Filter by Publication Year
Use the publication year filter to access established research, which provides a solid foundation for hypothesis generation and data analysis, and also newer studies which uncover emerging trends and innovative approaches.
Filter by Clinical Trial Count
By filtering for clinical trials, you can pinpoint genes and variants that are being actively researched and have promising therapeutic potential.
Filter by Disease Pleiotropy Index or Disease Specificity Index
Sort results by Disease Pleiotropy Index (DPI) and Disease Specificity Index (DSI) to refine your analysis using the REST API.
DPI measures the similarity or diversity of diseases associated with a gene or variant. A high DPI indicates that the associated diseases are closely related within a specific MeSH class, while a low DPI suggests a wider range of unrelated diseases.
DSI quantifies the breadth of a gene or variant’s disease associations. A high DSI indicates a gene or variant associated with a narrow set of diseases, while a low DSI suggests a broader range of associated diseases.

