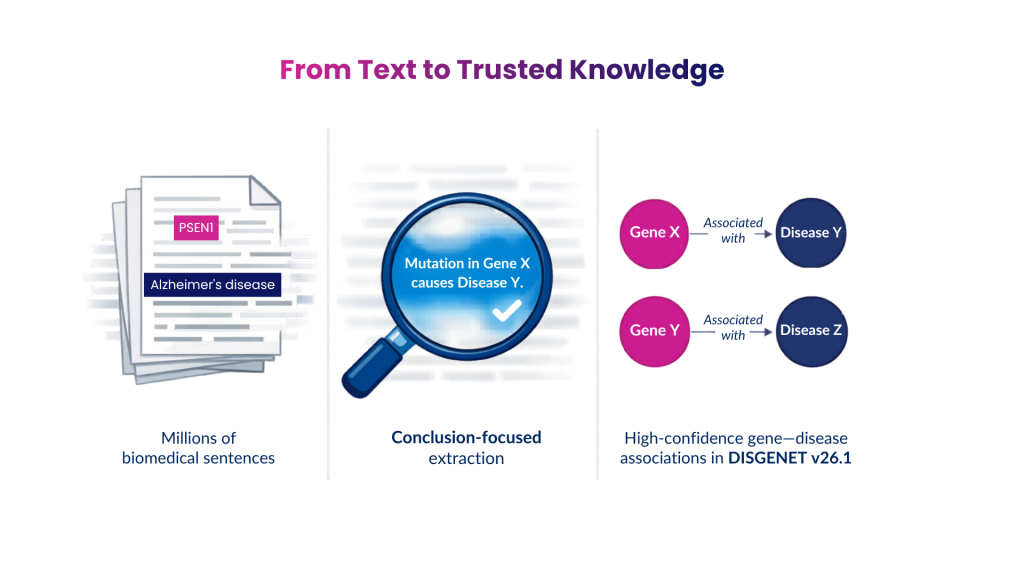
The latest update to DISGENET (v26.1) represents a significant step forward in extracting biomedical knowledge from scientific publications. This release is powered by an enhanced Relation Extraction (RE) model operating within a sophisticated, end-to-end NLP pipeline engineered for precision and scale. This system translates millions of biomedical publications into a structured network of gene/variant–disease relationships, streamlining the integration of current scientific knowledge into the DISGENET database.
What Is Relation Extraction?
Relation Extraction identifies meaningful relationships between entities mentioned in text. After detecting biomedical entities such as genes, variants, and diseases, the RE model determines whether those entities are biologically related within the sentence context.
For example, from
“Our findings demonstrate that loss-of-function mutations in Gene X drive progression of Disease Y”
the RE model extracts the structured association:
(Gene X) → positively associated with → (Disease Y)
These structured relationships allow the system to systematically capture and organize insights from vast amounts of biomedical literature.
Prioritizing Conclusion-Based Evidence
Accurately interpreting relationships in biomedical text requires more than recognizing co-occurrence: it demands an understanding of how entities interact and what kind of evidence supports that interaction. A key strength of the enhanced RE model is its ability to distinguish sentences that report experimental results from those focused on hypotheses or methodological descriptions.
By prioritizing statements grounded in empirical evidence, such as causal claims or statistically supported findings, the model ensures that gene/variant disease relationships reflect actionable biological insights rather than speculative or contextual mentions. This result-focused optimization significantly improves the relevance of the associations integrated into DISGENET v26.1, making the data more actionable for downstream research and decision-making.
A Hybrid Approach to Annotation
To train and validate this new RE model, DISGENET combines large language model (LLM)–assisted annotation with expert curation.
- LLMs rapidly scan millions of sentences, identifying potential relationships and linguistic patterns.
- Human experts then review and refine these candidate relations, resolving ambiguities and ensuring that the retained associations reflect true biological findings.
This synergy between automated scalability and expert precision ensures that DISGENET maintains broad coverage without compromising quality.
Why Accurate Literature Mining Matters
Extracting gene–disease and variant–disease associations underpins much of translational and precision medicine. Yet with millions of new biomedical publications released each year, manual curation alone cannot keep pace — critical findings risk being missed, delayed, or inconsistently captured. Moreover, selective curation can inadvertently introduce disease- or gene-specific biases, leaving gaps in coverage.
Automated extraction at scale addresses these challenges by ensuring comprehensive, unbiased coverage across the entire biomedical literature. However, doing this correctly is far from trivial: scientific text is rife with ambiguities, negations, context-dependent claims, and complex multi-entity statements that demand sophisticated interpretation.
By leveraging this new RE model, DISGENET v26.1 delivers clearer, contextually robust associations while maintaining broad, unbiased coverage. These data empower researchers to:
- Identify genes involved in disease mechanisms
- Explore genetic risk factors and biomarkers
- Support drug discovery and target prioritization
- Interpret genomic variants in clinical contexts
The result is a high-precision, literature-derived knowledge that transforms unstructured biomedical text into actionable insights, advancing biomedical discovery at a scale that manual methods simply cannot achieve.
