Full Traceability: Know Exactly Where Every Association Comes From
In research and drug discovery, the source of an association matters as much as the association itself. Provenance — the ability to trace each gene–disease or variant–disease association back to its original database, publication, or evidence type — is what transforms a data point into a trusted, decision-ready signal.
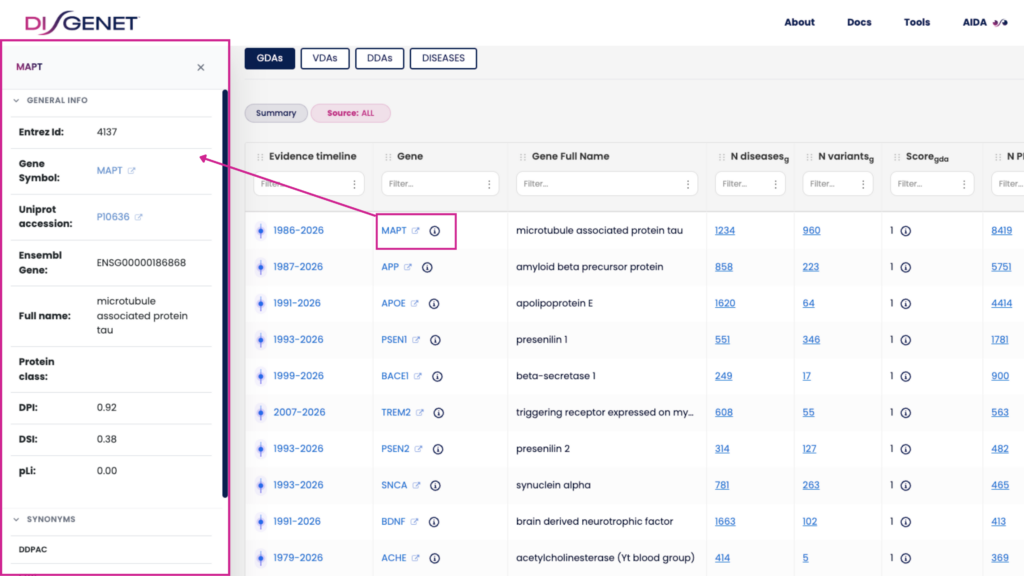
In pharma R&D, provenance enables target teams to assess whether a candidate is supported by specialized curated databases, preclinical models, or clinical studies — information that is critical at every triage gate. In biomarker discovery, it helps researchers distinguish well-established signals from emerging ones. In clinical genomics, it supports interpretability: can you point to the primary source, evaluate its quality, and defend your interpretation? With v26.1, the answer is a clearer yes.
A smarter NLP pipeline: result-focused evidence only
Not all sentences in a scientific paper carry the same evidential weight. A statement of hypothesis is not an experimental result. A method’s description is not a finding. The new Relation Extraction (RE) model in v26.1 understands this distinction: it filters for sentences reporting experimental outcomes and excludes those focused on hypotheses, methods, or background. This result-focused optimization directly improves the relevance and actionability of NLP-derived associations in DISGENET v26.1. For a deeper look at the model, read the dedicated blog post →
Supporting evidence is further classified using the new Polarity attribute, enabling users to differentiate between positive and negative findings directly within the Evidence table. This enables a more nuanced interpretation of gene–disease and variant–disease relationships, facilitating a more comprehensive assessment of consistency and conflicting evidence across studies.
New publication attributes: impact and translation potential
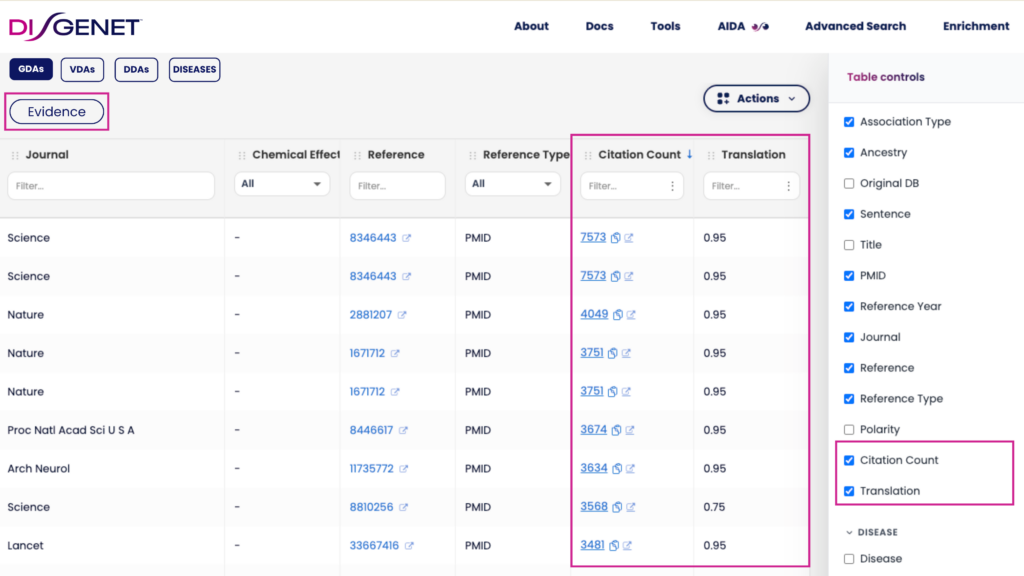
Each publication supporting an association now displays two additional attributes:
- Citing article count — the number of articles that cite the publication, with links to those citing works, so you can immediately assess its scientific influence and follow the evidence trail forward in time.
- Translation — a new metric reflecting the translational potential of each publication, from bench to bedside. This allows users to weigh evidence not only by volume or recency, but by its proximity to clinical application.
Together, these attributes provide a richer context for every evidence item, supporting more nuanced judgments about data quality and clinical relevance.
Prioritize with Confidence: Score Inspection, Enrichment, and Temporal Analysis
Explore what drives each score
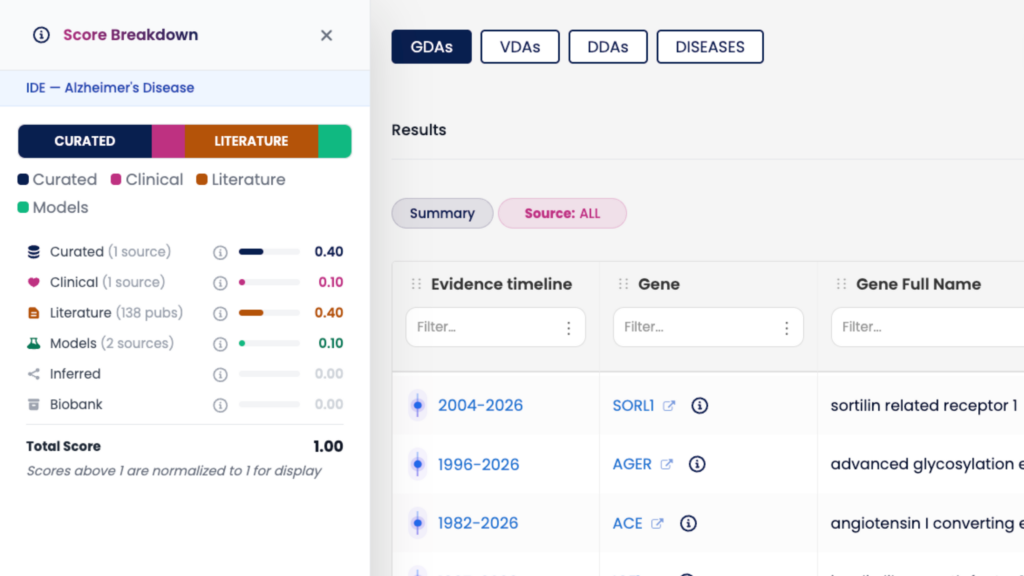
The DISGENET score integrates multiple evidence types into a single ranking signal. In v26.1, you can now explore the contribution of each evidence component directly — click on any score value in the interface to inspect the underlying factors and understand what is driving it. This capability is available both in the web interface and via the REST API.
This level of transparency matters for target prioritization: instead of treating the score as a black box, you can assess whether a high-scoring GDA is driven by curated database support, clinical evidence, NLP-extracted findings, or a combination — and weight your decisions accordingly.
Visualize how evidence has evolved over time
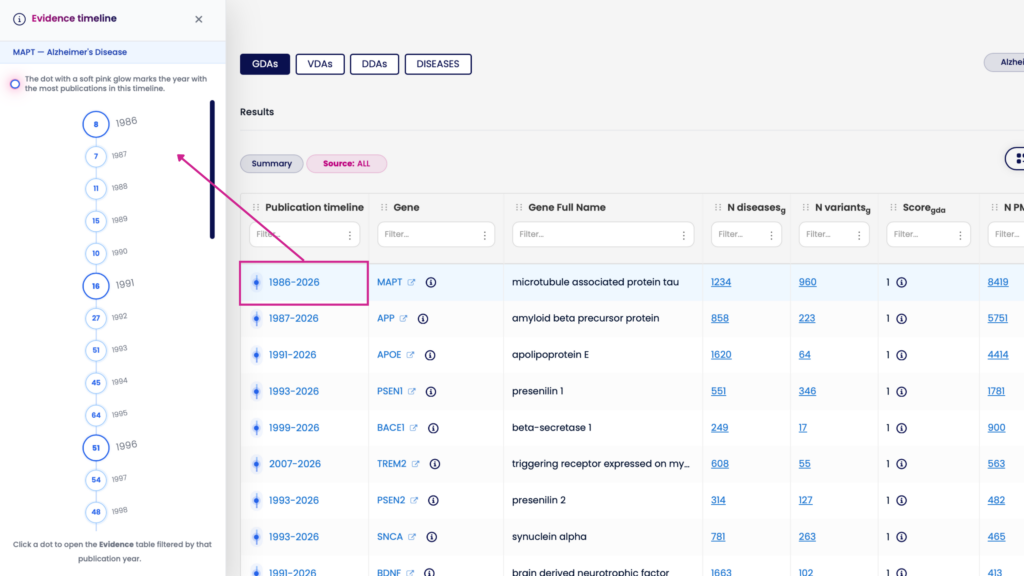
For every association, the web interface now includes a timeline visualization showing how the supporting evidence has grown, changed, or shifted over time. This is particularly valuable for:
- Target validation: Is this association a recent discovery or has it been accumulating evidence for years?
- Biomarker discovery: Is the signal gaining momentum or plateauing?
- Clinical genomics: Has the clinical relevance of this variant been consistently reported, or does it fluctuate?
The R package has been extended with new temporal analysis functions, enabling users to run this type of time-series analysis programmatically on their gene or variant for a particular disease.
Disease Enrichment Analysis — now fully integrated into your search workflow
Disease Enrichment Analysis can now be launched directly from selected gene or variant sets within your search results — no need to navigate away or reformat your data. The analysis now includes:
- Odds ratio metrics for more robust statistical interpretation
- Customizable plots that can be adapted to your reporting needs
- Downloadable outputs for downstream analysis and publication
Improved Advanced Search
Advanced Search has been enhanced with more flexible text matching — handling synonyms, partial terms, and variant nomenclature more reliably — and now returns results ranked by relevance rather than by a fixed default order. Results can be downloaded for further analysis.
Meet the New AIDA
AIDA — the DISGENET AI assistant — has been rebuilt on an updated architecture and significantly expanded in scope. In addition to genes, variants, and diseases, AIDA now covers therapies, compounds, and disease subtypes, making it possible to ask cross-entity questions that previously required manual multi-step queries.
Whether you are running a target landscape assessment, exploring drug repurposing hypotheses, or mapping the evidence landscape for a disease subtype, AIDA provides a natural-language entry point into the DISGENET knowledge graph — grounded in curated, traceable evidence.
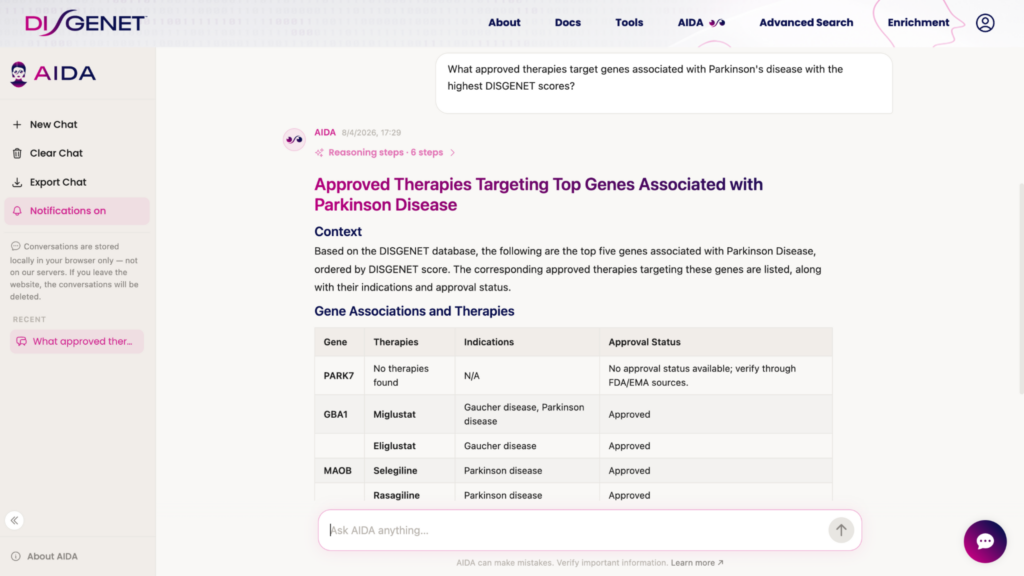
A Redesigned Interface Built Around Your Workflow
The DISGENET web interface has been substantially updated in v26.1, with every change aimed at reducing friction and accelerating insight.
Navigation and orientation:
- A redesigned navigation bar and breadcrumb navigation let you track your exploration path and move between related entities without losing context.
- New side panels surface contextual information on entities and scoring without requiring a page change.
Working with data:
- Tables are now more interactive, with customizable layouts, built-in filtering, and one-click access to features such as Favorites and MyScore.
- Action buttons for enrichment, saving, and downloading are now directly accessible from the results view.
- The score component explorer (described above) is embedded directly into the table, enabling inline evidence inspection.
Documentation:
- The Docs section has been updated with more visual explanations of metrics and scoring.
- A new dedicated section on Disease Enrichment Analysis is now available.
Whether you are exploring targets in pharma R&D, validating biomarkers, or interpreting variants in a clinical context, the updated interface is designed to help you move faster — from initial search to actionable conclusion.
Try It Now
DISGENET v26.1 is live. Log in to explore the new provenance attributes, run your Disease Enrichment Analysis on the updated interface, inspect the score components for your targets, or ask AIDA a question about your disease area.
